Biopython
Introduction
From the official Biopython project website:
Biopython is a set of freely available tools for biological computation written in Python by an international team of developers.
It is a distributed collaborative effort to develop Python libraries and applications which address the needs of current and future work in bioinformatics. The source code is made available under the Biopython License, which is extremely liberal and compatible with almost every license in the world.
This workshop assumes a working knowledge of the Python programming language and basic understanding of the concepts of online DNA and Protein sequence repositories.
Introductions to Python can be found here and here.
Getting Started
Python code examples
The Python scripts and data files for this workshop can be downloaded from here. On your computer, unzip the downloaded folder and use it as working directory for this workshop.
Python programming environment
The Anaconda environment from Anaconda Inc. is widely used because it bundles a Python interpreter, most of the popular packages, and development environments. It is cross platform and freely available. There are two somewhat incompatible versions of Python; version 2.7 is deprecated but still fairly widely used. Version 3 is the supported version.
Note: The latest Biopython package version (1.77+) requires Python 3.
-
Visit the Anaconda download website and download the installer for Python 3 for your operating system (Windows, Mac OSX, or Linux). We recommend to use the graphical installer for ease of use.
-
Launch the downloaded installer, follow the onscreen prompts and install the Anaconda distribution on your local hard drive.
The
Anaconda Documentation provides an introduction to the Ananconda environment and bundled applications. For the purpose of this workshop we focus on the Anaconda Navigator and Spyder.
Navigator
Once you have installed Anaconda, start the Navigator application:
You should see a workspace similar to the screenshot, with several options for working environments, some of which are not installed. We will use Spyder which should already be installed. If not, click the button to install the package.
Spyder
Now we will switch to Spyder. Spyder is an Integrated Development Environment, or IDE, aimed at Python. It is well suited for developing longer, more modular programs.
- To start it, return to the
Anaconda Navigatorand click on theSpydertile. It may take a while to open (watch the lower left of the Navigator). - Once it starts, you will see a layout with an editor pane on the left, an explorer pane at the top right, and an iPython console on the lower right. This arrangement can be customized but we will use the default for our examples. Type code into the editor. The explorer window can show files, variable values, and other useful information. The iPython console is a frontend to the Python interpreter itself. It is comparable to a cell in JupyterLab.
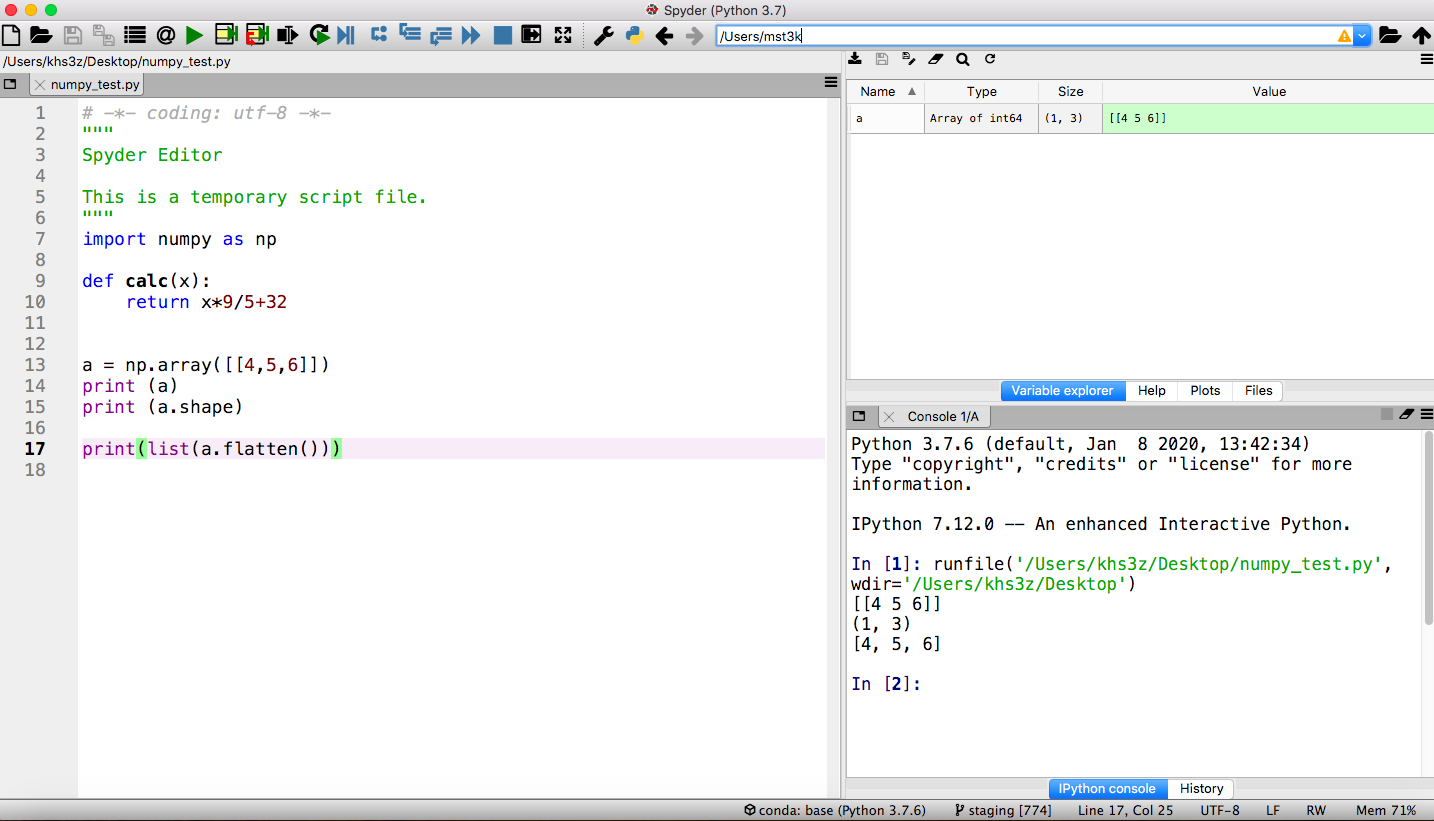
Installation of the Biopython package
It is recommended to install the biopython package from PyPI using the pip install command. Detailed instructions are available
here.
On your own computer:
Start the Anaconda Prompt command line tool following the instructions for your operating system.
- Start Anaconda Prompt on Windows
- Start Anaconda Prompt on Mac, or open a terminal window.
- Linux: Just open a terminal window.
At the prompt, type the following command and press enter/return:
pip install biopython
This command will install the latest biopython package version in your current Anaconda Python environment.
On Rivanna (UVA’s HPC platform):
Rivanna offers several Anaconda distributions with different Python versions. Before you use Python you need to load one of the anaconda software modules and then run the pip install command.
module load anaconda
pip install --user biopython
Note: You have to use the --user flag which instructs the interpreter to install the package in your home directory. Alternativley, create your own custom
Conda environment first and run the pip install biopython command in that environment (without --user flag)
Testing the Biopython Installation
Start the Spyder IDE (see
here). In the IPython console pane, type the following command and press enter/return:
import Bio
print (Bio.__version__)
If the package is installed correctly, the output will show the biopython version number.
Bio Subpackages and Classes
The Bio package provides a large number of subpackages providing specific functionality. The Biopython website provides a
full list of all subpackages.
The following table shows an excerpt of that list relevant for this workshop.
| Subpackages/Classes | Purpose |
|---|---|
| Bio.Entrez | Functions to retrieve Entrez records and associated data |
| Bio.ExPASy | Tools to access data hosted on the ExPASy protein databases |
| Bio.SwissProt | Tools to work with the sprotXX.dat file from SwissProt |
| Bio.Seq | Sequence datastructure (immutable=read-only) |
| Bio.MutableSeq | Sequence datastructure (mutable=modifiable |
| Bio.SeqRecord | Datastucture for Seq object plus enriched information |
| Bio.SeqIO | Read/write sequences (various file formats ) |
| Bio.AlignIO | A new multiple sequence Alignment Input/Output interface for BioPython 1.46 and later |
| Bio.Align.MultipleSeqAlignment | Tools for Code for working with sequence alignments |
Online Datasets and Databases
The Bio module provides several classes to process output and datasets provided by the following web services and tools:
- FASTA
- Blast output – both from standalone and WWW Blast
- Clustalw
- GenBank
- PubMed and Medline
- ExPASy files, like Enzyme and Prosite
- SCOP, including ‘dom’ and ‘lin’ files
- UniGene
- SwissProt
In this workshop we will explore options to download nucleotide and protein sequences from Entrez and ExPASy.
Accessing NCBI’s Entrez Databases
The Bio.Entrez submodule provides access to the Entrez databases. When you use this module you need to know the String descriptor of the database you want to query (aka its name). A list of valid database names is provided in
column three of this table.
Note: Please review the General Introduction to the E-utilities for accessing the Entrez Application Programming Interface Programm. The E-utilities limit the frequency of API calls and your IP address may be blocked if you contineously exceed the limit.
Basic Steps:
- Provide an email address. This is required!
Entrez.email = "your@somewhere". - Use an
E-utility to get a handle for the data of interest, e.g.
handle = Entrez.esearch(...). - Use handle to read or parse data with
handle.read()orhandle.parse(). - Close the handle with
handle.close().
Also read “What the heck is a handle?"
Find Database Records:
Let’s find the protein records associated with the human Pax6 gene and download the associated sequences in FASTA format.
To search the database we use the Entrez.esearch function. We need to specify the database via the db, argument and specify a search term (provided as a list of Strings).
The following code is available in the entrez-fasta.py file.
from Bio import Entrez
Entrez.email = "YOU@SOMEWHERE.com" # your email address is required
handle = Entrez.esearch(db="protein", term = ["Homo sapiens[Orgn] AND pax6[Gene]"], usehistory="y")
record = Entrez.read(handle)
handle.close()
# iterate over items
for k,v in record.items():
print (k,v)
Output
Count 88
RetMax 20
RetStart 0
QueryKey 1
WebEnv MCID_5f87b7433f6876111801786c
IdList ['1587062735', '1587062729', '1587062727', '1587062721', '1587062719', '1587062717', '1587062715', '1587062713', '1587062710', '1587062708', '1587062706', '1587062703', '1587062701', '1587062699', '1587062697', '1587062695', '1587062690', '1587062688', '1587062686', '1587062684']
TranslationSet [{'From': 'Homo sapiens[Orgn]', 'To': '"Homo sapiens"[Organism]'}]
TranslationStack [{'Term': '"Homo sapiens"[Organism]', 'Field': 'Organism', 'Count': '1423829', 'Explode': 'Y'}, {'Term': 'pax6[Gene]', 'Field': 'Gene', 'Count': '2621', 'Explode': 'N'}, 'AND']
QueryTranslation "Homo sapiens"[Organism] AND pax6[Gene]
The search results are returned as a
dictionary and we can retrieve the list of unique IDs that match our query via record["IdList"].
Note: The IdList returned by esearch is limited to the top 20 hits by default (defined by retmax). There are two workarounds:
- Use the
retmax=<number>keyword argument to increase the maximum number of retrieved records. The problem is you need to know what a reasonable number is. - Or better, use the
usehistory='y'keyword argument. This will save the search results on the remote server and provideWebEnvandQueryKeyentries that can be used with theefetchfunction (see next section) to retrieve all search records (beyond the top 20).
By default the returned IDs reflect the GI numbers. The accession.version numbers can be retrieved instead by passing idtype='acc' as an optional keyword argument to the esearch function. See the
detailed documentation of the esearch function here.
Download and save sequences as FASTA file:
With the ID list in hand, we can now download the sequence records using just a few lines of code and save them in a single multi-sequence FASTA file. The efetch function is used when you want to retrieve a full record from Entrez.
# fetch records using id list
handle = Entrez.efetch(db="protein", rettype="fasta", retmode="text", id=record["IdList"])
result = handle.read() # return type is simple string
handle.close()
# remove empty lines
fastaseq = result.replace("\n\n","\n")
with open('HsPax6-protein.fasta', 'w') as f:
f.write(fastaseq)
Alternatively, we can pull individual sequences one at a time and save each sequence into a separate file. To do this we implement a
for loop that iterates over this list and use the Entrez.efetch function to retrieve the FASTA sequence record associated with each id. We wrap this for loop in a
open file operation block to save the retrieved FASTA records into a single .fasta text file.
Let’s retrieve the nucleotide sequences of our previous top 5 ID hits as GenBank files. We specify the database with the db="nucleotide" and format with the rettype="gb" keyword arguments.
The code is provided in the entrez-genbank.py file.
from Bio import Entrez
Entrez.email = "YOU@SOMEWHERE.com" # provide your email address
handle = Entrez.esearch(db="nucleotide", term = ["Homo sapiens[Orgn] AND pax6[Gene]"], retmax=5, usehistory="y")
record = Entrez.read(handle)
handle.close()
for k,v in record.items():
print (k,v)
# iterate over ids in list
for seq_id in record["IdList"]:
# get entry from Entrez
print (seq_id)
handle = Entrez.efetch(db="nucleotide", id=seq_id, rettype="gb", retmode="text")
result = handle.read()
handle.close()
# save
filename = f"HsPax6-{seq_id}-nucleotide.gb"
print ("Saving to file:", filename)
with open(filename, 'w') as gbfile:
# append fasta entry
gbfile.write(result.rstrip()+"\n")
Output:
Count 106
RetMax 5
RetStart 0
QueryKey 1
WebEnv MCID_5f87bb652dd862297e1727ea
IdList ['1844161464', '1844139642', '1844139635', '1844139631', '1844139629']
TranslationSet [{'From': 'Homo sapiens[Orgn]', 'To': '"Homo sapiens"[Organism]'}]
TranslationStack [{'Term': '"Homo sapiens"[Organism]', 'Field': 'Organism', 'Count': '27682287', 'Explode': 'Y'}, {'Term': 'pax6[Gene]', 'Field': 'Gene', 'Count': '3232', 'Explode': 'N'}, 'AND']
QueryTranslation "Homo sapiens"[Organism] AND pax6[Gene]
1844161464
Saving to file: HsPax6-1844161464-nucleotide.gb
1844139642
Saving to file: HsPax6-1844139642-nucleotide.gb
1844139635
Saving to file: HsPax6-1844139635-nucleotide.gb
1844139631
Saving to file: HsPax6-1844139631-nucleotide.gb
1844139629
Saving to file: HsPax6-1844139629-nucleotide.gb
Done
Note that the record['IdList'] may not represent all the records. Remember that the record['WebEnv'] and record['QueryKey'] entries provide access to the search history on the remote server. So we can use these instead of the record['IdList'] to get all records.
# Alternative: fetch all records using search history
handle = Entrez.efetch(db="protein", rettype="fasta", retmode="text", webenv=record["WebEnv"], query_key=record["QueryKey"])
Exercise: Find and download the top 10 FASTA EST nucleotide sequences for the mouse (Mus Musculus) TP53 tumor suppressor. Hint: look up the EST database descriptor in this table.
Solution:
handle = Entrez.esearch(db="nucest", term = ["Mus musculus[Orgn] AND tp53[Gene]"], retmax=10, usehistory="y")
record = Entrez.read(handle)
handle.close()
for k,v in record.items():
print (k,v)
# iterate over ids in list
for seq_id in record["IdList"]:
# get entry from Entrez
print (seq_id)
handle = Entrez.efetch(db="nucest", id=seq_id, rettype="fasta", retmode="text")
result = handle.read()
handle.close()
# save
filename = f"MmP53-{seq_id}-est.fasta"
print ("Saving to file:", filename)
with open(filename, 'w') as fastafile:
# append fasta entry
fastafile.write(result.rstrip()+"\n")
Retrieve Protein Records from the ExPASy Database
Here are a few examples demonstrating how to access the ExPASy databases Swissport and Prosite. The Biopython documentation provides more details.
Swiss-Prot
from Bio import ExPASy
from Bio import SwissProt
# get single protein record
accession_no = "O23729"
handle = ExPASy.get_sprot_raw(accession_no)
record = SwissProt.read(handle)
print (record.entry_name)
print (record.sequence_length)
print (record.data_class)
print (record.accessions)
print (record.organism)
# print first 10 aa
print (record.sequence[:10]) # string
Output:
CHS3_BROFI
394
Reviewed
['O23729']
Bromheadia finlaysoniana (Orchid).
MAPAMEEIRQ
The return type of SwissProt.read() is a
Bio.SwissProt.Record object. In the above example we’re printing only a subset of its fields. The record.sequence field is a string, but it can easily be converted into a
Bio.Seq object.
Tip: Use dir(record) to get a list of all record attribute names.
Prosite
from Bio import ExPASy
from Bio.ExPASy import Prosite
handle = ExPASy.get_prosite_raw("PS00001")
record = Prosite.read(handle)
print (record.name)
print (record.type) # e.g. PATTERN, MATRIX, or RULE
print (record.pattern)
print (record.rules)
print (record.matrix)
Output:
ASN_GLYCOSYLATION
PATTERN
N-{P}-[ST]-{P}.
[]
[]
The return type of Prosite.read() is a
Bio.ExPASy.Prosite.Record object.
Note: Use the Bio.ExPASy.Prosite.parse() function to parse files containing multiple records.
Prosite Documentation
from Bio import ExPASy
from Bio.ExPASy import Prodoc
handle = ExPASy.get_prosite_raw("PDOC00001")
record = Prodoc.read(handle)
Exercise: Retrieve the SwissProt records for proteins with the following IDs: “O23729”, “O23730”, “O23731”. Try to use list comprehension to create a list containing the records for all retrieved proteins.
Solution
from Bio import ExPASy,SwissProt
accession_nos = ["O23729", "O23730", "O23731"]
handles = [ExPASy.get_sprot_raw(a) for a in accession_nos]
records = [SwissProt.read(h) for h in handles]
for record in records:
print(record.entry_name)
print(",".join(record.accessions))
print(record.keywords)
print(repr(record.organism))
print(record.sequence[:20],"\n")
Learn more about SwissProt.
ScanProsite
We can query the Prosite database with protein sequences or motifs to find proteins with corresponding matches, see ScanProsite for details.
Option 1: Submit protein sequence (use the seq= keyword argument)
- UniProtKB accessions e.g. P98073
- Identifiers e.g. ENTK_HUMAN
- PDB identifiers e.g. 4DGJ
- Sequences in FASTA format.
Option 2: Submit motif sequence (use the sig= keyword argument)
- PROSITE accession e.g. PS50240
- Identifier e.g. TRYPSIN_DOM
- Your own pattern e.g. P-x(2)-G-E-S-G(2)-[AS].
- Combinations of motifs can also be used.
from Bio.ExPASy import ScanProsite
uniprot_id = "P26367" # human Pax-6
handle = ScanProsite.scan(seq=uniprot_id)
results = ScanProsite.read(handle)
By executing handle.read(), you can obtain the search results in raw XML format. Here we use Bio.ExPASy.ScanProsite.read to parse the raw XML into a Bio.ExPASy.ScanProsite.Record object which represents a specialized list.
We can now access the found matches like this:
print ("Number of matches:", results.n_match)
for r in results:
print (r)
Output:
Number of matches: 4
{'sequence_ac': 'P26367', 'sequence_id': 'PAX6_HUMAN', 'sequence_db': 'sp', 'start': 4, 'stop': 130, 'signature_ac': 'PS51057', 'signature_id': 'PAIRED_2', 'score': '64.941', 'level': '0'}
{'sequence_ac': 'P26367', 'sequence_id': 'PAX6_HUMAN', 'sequence_db': 'sp', 'start': 38, 'stop': 54, 'signature_ac': 'PS00034', 'signature_id': 'PAIRED_1', 'level_tag': '(0)'}
{'sequence_ac': 'P26367', 'sequence_id': 'PAX6_HUMAN', 'sequence_db': 'sp', 'start': 208, 'stop': 268, 'signature_ac': 'PS50071', 'signature_id': 'HOMEOBOX_2', 'score': '20.164', 'level': '0'}
{'sequence_ac': 'P26367', 'sequence_id': 'PAX6_HUMAN', 'sequence_db': 'sp', 'start': 243, 'stop': 266, 'signature_ac': 'PS00027', 'signature_id': 'HOMEOBOX_1', 'level_tag': '(0)'}
You see that each item r represents a dictionary describing a specific match.
Working with Sequence Files
Sequence Objects
The Seq object is similar to a string object augmented with methods for nucleotide sequence operations including
find(),count()complement(),reverse_complement()transcribe(),back_transcribe()translate()
The following code examples are in the seq.py script.
from Bio.Seq import Seq
my_dna = Seq("ATGAGTACACTATAGA")
print (my_dna)
# find position of first subsequence
print (my_dna.find("TAC"))
print (my_dna.find("AC"))
print (my_dna.find("CTC"))
# count
print ("A count:", my_dna.count("A"))
print ("AC count:", my_dna.count("AC"))
print ("AA count:", Seq("AAAA").count("AA")) # non-overlapping
Output:
ATGAGTACACTATAGA
5
6
-1
A count: 7
AC count: 2
AA count: 2
Note the return of -1 if no sequence match was found.
# complement and reverse complement
compl = my_dna.complement()
rev_compl = my_dna.reverse_complement()
print ("original: \t", my_dna)
print ("complement:\t", compl)
print ("rev complement:\t", rev_compl)
# transcription
my_rna = my_dna.transcribe()
print ("RNA:", my_rna)
# translation
my_peptide = my_dna.translate()
print ("Peptide:", my_peptide)
Output:
original: ATGAGTACACTATAGA
complement: TACTCATGTGATATCT
rev complement: TCTATAGTGTACTCAT
RNA: AUGAGUACACUAUAGA
Peptide: MSTL*
Like Strings, Seq objects are immutable; this means that the sequence is read-only and cannot be modified in place. However, you can convert a Seq object into a MutableSeq object that allows you to manipulate he sequence after object initialization.
my_dna[2] = 'A' # error, immutable Seq object
mutable_dna = my_dna.tomutable()
mutable_dna[2] = 'A'
print (my_dna)
print (mutable_dna)
Output:
ATGAGTACACTATAGA
ATAAGTACACTATAGA
Note that the sequence is zero-indexed: the first nucleotide has index 0, the second has index 1, and so forth. So in this example we’re changing the third nucleotide (index 2, G->A).
Exercise: Create a Seq object with a DNA nucleotide sequence of your choice. Find the first putative start codon (ATG), replace each “C” with a “G”, and transcribe and translate the original as well as the modified sequence. Hint: As an intermediary step, convert Seq object to a string and use a string method for replacement.
Solution:
from Bio.Seq import Seq
seq = Seq("CTGACTGGATGACCATTGGGCAACTACCCATGACTAGTTAAGTAATTTTTAAAAA")
atg_pos = seq.find("ATG")
cds = seq[atg_pos:]
peptide = cds.translate(to_stop=True)
mod_seq = Seq(str(seq).replace("C","G"))
mod_cds = mod_seq[atg_pos:]
mod_peptide = mod_cds.translate(to_stop=True)
print ("DNA (original):", seq)
print ("DNA (modified):", mod_seq)
print ("Peptide (original):", peptide)
print ("Peptide (modified):", mod_peptide)
Handling Sequence Records
The SeqRecord class provides the following fields:
.seq: a sequence (as a Seq object).id: the identifier, e.g. an accession number (String).name: can be just the accession number or the locus name (String).description: self-explanatory (String).annotations: dictionary of additional often unstructured info (optional).letter_annotations: often used for quality scores or secondary structure info.features: list of SeqFeature objects; more structured than annotations, e.g. gene position in a genome, or domain position in a protein.dbxref: list of database cross-references
So it is used to wrap around a
Seq object with richer information. We can manually create a SeqRecord object like this:
from Bio.Seq import Seq
from Bio.SeqRecord import SeqRecord
record = SeqRecord(
Seq("MKQHKAMIVALIVICITAVVAALVTRKDLCEVHIRTGQTEVAVF"),
id="YP_025292.1",
name="HokC",
description="toxic membrane protein, small")
print(record)
The above code example is in the seqrecord.py script.
Output:
ID: YP_025292.1
Name: HokC
Description: toxic membrane protein, small
Number of features: 0
Seq('MKQHKAMIVALIVICITAVVAALVTRKDLCEVHIRTGQTEVAVF')
Sequence File Operations
The Bio.SeqIO class provides simple tools to read and write a variety of sequence file formats (including multiple sequence alignments). It operates exclusively on SeqRecord objects.
Read Fasta File
from Bio import SeqIO
file = open('HsPax6-protein.fasta')
fastarecords = SeqIO.parse(file, "fasta")
# create a list of SeqRecords
fastalist = [entry for entry in fastarecords]
# iterate over fasta entries
for entry in fastalist:
print (f"ID={entry.id}")
print (f"Name={entry.name}")
print (f"Description={entry.description}")
print (f"Seq length={len(entry.seq)}")
print (f"Features={entry.features}") # empty for Fasta format
print (f"Sequence={entry.seq}\n")
Exercise: Filter the list of records to only include sequences with less than 300 amino acids.
Solution
# filter list of records
sublist = [e for e in fastalist if len(e.seq) < 300]
print (f"Total number of sequences: {len(fastalist)}")
print (f"Number of sequences (<300 aa): {len(sublist)}")
Convert Genbank to Fasta File
from Bio import SeqIO
gb_file = 'HsPax6-208879460-nucleotide.gb'
with open(gb_file) as f:
gb_generator = SeqIO.parse(f, format='gb')
for entry in gb_generator:
with open(f'Hs-pax6-{entry.id}-nucleotide.fasta', 'w') as fileout:
SeqIO.write(entry, fileout, 'fasta')
The later versions of Biopython also include a Bio.SeqIO.convert() function.
# convert GenBank to Fasta
count = Bio.SeqIO.convert("my_file.gb", "genbank", "my_file.fasta", "fasta")
AlignIO: Reading Sequence Alignment Files
The
Bio.AlignIO class provides functions to handle paired or multiple sequence alignment files. It does not perform the alignment but provides tools to read/write alignment files and manipulate alignment objects. Bio.AlignIO uses the same set of functions for input and output as in Bio.SeqIO, and the same names are supported for the file formats.
The key functions are:
Bio.AlignIO.read(): For a file that contains one and only one alignment. The return type is aBio.Align.MultipleSeqAlignmentobject.Bio.AlignIO.parse(): A more general function when the file may contain multiple separate alignments. The return type is a generator that can be converted into a list ofBio.Align.MultipleSeqAlignmentobjects.
Example:
Let’s create a Fasta file with Pax6 orthologs from human, mouse, xenopus, pufferfish, zebrafish (2), and fruitfly. The following code example is in the createPax6_fasta.py script.
from Bio import Entrez
# get human, mouse, xenopus, pufferfish, zebrafish 1, zebrafish 2, Drosophila 1, Drosophila 2
ids = ['1587062735','AAH36957.1','NP_001006763.1','XP_029701655.1','NP_571379.1','NP_571716.1','NP_524628','NP_524638']
handle = Entrez.efetch(db="protein", rettype="fasta", retmode="text", id=ids)
result = handle.read()
handle.close()
fastaseq = result.replace("\n\n","\n")
with open('Pax6-multispec-protein.fasta', 'w') as f:
f.write(fastaseq)
This will create the Pax6-multispec-protein.fasta Fasta file with 8 sequences. The alignment was performed using
Clustal Omega and you can
download the Pax6-multispec-protein.aln alignment file and move it to your Python script folder that you use for this workshop.
Alternatively, create the alignment yourself:
- Visit the
Clustal Omega website and upload the
Pax6-multispec-protein.fastafile as input. - Under Step 1, click the Choose File button and upload the
Pax6-multispec-protein.fastafile as input. - Under Step 3, click Submit.
- When the alignment is done, click the Alignments tab, select the entire alignment output in the window and paste it into a text editor. Do not use Microsoft Word for this but programs like
Text Edit,Notepad++,Emacsorviminstead. - Save the alignment in the text editor as
Pax6-multispec-protein.alnin your Python script folder that you use for this workshop.
The following code examples are in the alignio-parse_clustal.py script.
Parse the alignment file
from Bio import AlignIO
inputfile = open("Pax6-multispec-protein.aln", "r")
# assuming single alignment in file; use AlignIO.parse for multiple alignments
alignment = AlignIO.read(inputfile, "clustal")
inputfile.close()
print ("Alignment length:", alignment.get_alignment_length())
print (alignment,"\n")
Output:
Alignment with 8 rows and 867 columns
MFTLQPTPTAIGTVVPPWSAGTLIERLPSLEDMAHKGHSGVNQL...PWV NP_524628.2
---MMLTTEHIMHGHPH-----SSVGQSTLFGCSTAGHSGINQL...--- NP_524638.3
---------------------------------MQNSHSGVNQL...--- NP_001006763.1
--------------------------------------------...--- NP_001355831.1
---------------------------------MQNSHSGVNQL...--- AAH36957.1
--------------------------------MMQNSHSGVNQL...--- XP_029701655.1
----MPQKEY-Y----N-----RATWESGVASMMQNSHSGVNQL...--- NP_571379.1
----MPQKEY-H----N-----QPTWESGVASMMQNSHSGVNQL...--- NP_571716.1
Update identifier
species = ['H.sapiens', 'M.musculus', 'X.tropicalis', 'T.rubripes', 'D.rerio', 'D.rerio', 'D.melanogaster', 'D.melanogaster']
for idx,line in enumerate(alignment):
line.id = f"{species[idx]}:{line.id}"
print (alignment)
Output:
Alignment with 8 rows and 867 columns
MFTLQPTPTAIGTVVPPWSAGTLIERLPSLEDMAHKGHSGVNQL...PWV H.sapiens:NP_524628.2
---MMLTTEHIMHGHPH-----SSVGQSTLFGCSTAGHSGINQL...--- M.musculus:NP_524638.3
---------------------------------MQNSHSGVNQL...--- X.tropicalis:NP_001006763.1
--------------------------------------------...--- T.rubripes:NP_001355831.1
---------------------------------MQNSHSGVNQL...--- D.rerio:AAH36957.1
--------------------------------MMQNSHSGVNQL...--- D.rerio:XP_029701655.1
----MPQKEY-Y----N-----RATWESGVASMMQNSHSGVNQL...--- D.melanogaster:NP_571379.1
----MPQKEY-H----N-----QPTWESGVASMMQNSHSGVNQL...--- D.melanogaster:NP_571716.1
Slicing and joining
# slice: first axis defines line, second axis defines column index (zero-indexed)
# get lines 1-6, first 50 columns
subset = alignment[:6,:50]
print (subset)
Output:
Alignment with 6 rows and 50 columns
MFTLQPTPTAIGTVVPPWSAGTLIERLPSLEDMAHKGHSGVNQLGGVFVG H.sapiens:NP_524628.2
---MMLTTEHIMHGHPH-----SSVGQSTLFGCSTAGHSGINQLGGVYVN M.musculus:NP_524638.3
---------------------------------MQNSHSGVNQLGGVFVN X.tropicalis:NP_001006763.1
-------------------------------------------------- T.rubripes:NP_001355831.1
---------------------------------MQNSHSGVNQLGGVFVN D.rerio:AAH36957.1
--------------------------------MMQNSHSGVNQLGGVFVN D.rerio:XP_029701655.1
Let’s join two alignment blocks:
edited = alignment[:,:50] + alignment[:,500:]
print (edited)
Output:
Alignment with 8 rows and 417 columns
MFTLQPTPTAIGTVVPPWSAGTLIERLPSLEDMAHKGHSGVNQL...PWV H.sapiens:NP_524628.2
---MMLTTEHIMHGHPH-----SSVGQSTLFGCSTAGHSGINQL...--- M.musculus:NP_524638.3
---------------------------------MQNSHSGVNQL...--- X.tropicalis:NP_001006763.1
--------------------------------------------...--- T.rubripes:NP_001355831.1
---------------------------------MQNSHSGVNQL...--- D.rerio:AAH36957.1
--------------------------------MMQNSHSGVNQL...--- D.rerio:XP_029701655.1
----MPQKEY-Y----N-----RATWESGVASMMQNSHSGVNQL...--- D.melanogaster:NP_571379.1
----MPQKEY-H----N-----QPTWESGVASMMQNSHSGVNQL...--- D.melanogaster:NP_571716.1
Exporting to other alignment file formats
# save as Stockholm
with open("Pax6-multispec-protein.sth", "w") as outputfile:
AlignIO.write(alignment, outputfile, "stockholm")
# get alignment as formatted string
print ("Formatted Alignment:")
print (format(alignment, "clustal"))
Output:
Formatted Alignment:
CLUSTAL X (1.81) multiple sequence alignment
H.sapiens:NP_524628.2 MFTLQPTPTAIGTVVPPWSAGTLIERLPSLEDMAHKGHSGVNQLGGVFVG
M.musculus:NP_524638.3 ---MMLTTEHIMHGHPH-----SSVGQSTLFGCSTAGHSGINQLGGVYVN
X.tropicalis:NP_001006763.1 ---------------------------------MQNSHSGVNQLGGVFVN
T.rubripes:NP_001355831.1 --------------------------------------------------
D.rerio:AAH36957.1 ---------------------------------MQNSHSGVNQLGGVFVN
D.rerio:XP_029701655.1 --------------------------------MMQNSHSGVNQLGGVFVN
D.melanogaster:NP_571379.1 ----MPQKEY-Y----N-----RATWESGVASMMQNSHSGVNQLGGVFVN
D.melanogaster:NP_571716.1 ----MPQKEY-H----N-----QPTWESGVASMMQNSHSGVNQLGGVFVN
Exercise: Find the first alignment block that shows no gaps across all 8 aligned sequences.
- Print the block.
- Save the block as a new clustal formatted text file.
- From that block, extract the D. rerio (zebrafish) sequences and print the two sequences
Solution:
from Bio import AlignIO
inputfile = open("Pax6-multispec-protein.aln", "r")
# assuming single alignment in file; use AlignIO.parse for multiple alignments
alignment = AlignIO.read(inputfile, "clustal")
inputfile.close()
length = alignment.get_alignment_length()
# create boolean list to indicate if all lines in column are wiithout gap
pattern = [all([a.seq[i] != "-" for a in alignment]) for i in range(length)]
print ("No gap in column:")
print (pattern,"\n")
# create list of column indices without gaps
full_indices = [i for i in range(length) if pattern[i]]
print ("Full column indices:")
print (full_indices,"\n")
# scan consecutive columns for completeness
firstblock_i = []
j=0
start=0
for i in full_indices[start:]:
if j==0 or i==full_indices[start]+j:
firstblock_i.append(i)
else:
break
j+=1
# slice alignment to show first block without gaps
block1 = alignment[:,firstblock_i[0]:firstblock_i[-1]+1]
print (block1)
# save
with open("Pax6-multispec-block1.aln", "w") as outputfile:
AlignIO.write(block1, outputfile, "clustal")
# get zebrafish sequence lines
zebrafish_lines = block1[4:6,:]
print (zebrafish_lines)